- #Osirix lite nifti to dicom for mac#
- #Osirix lite nifti to dicom software#
- #Osirix lite nifti to dicom code#
- #Osirix lite nifti to dicom series#
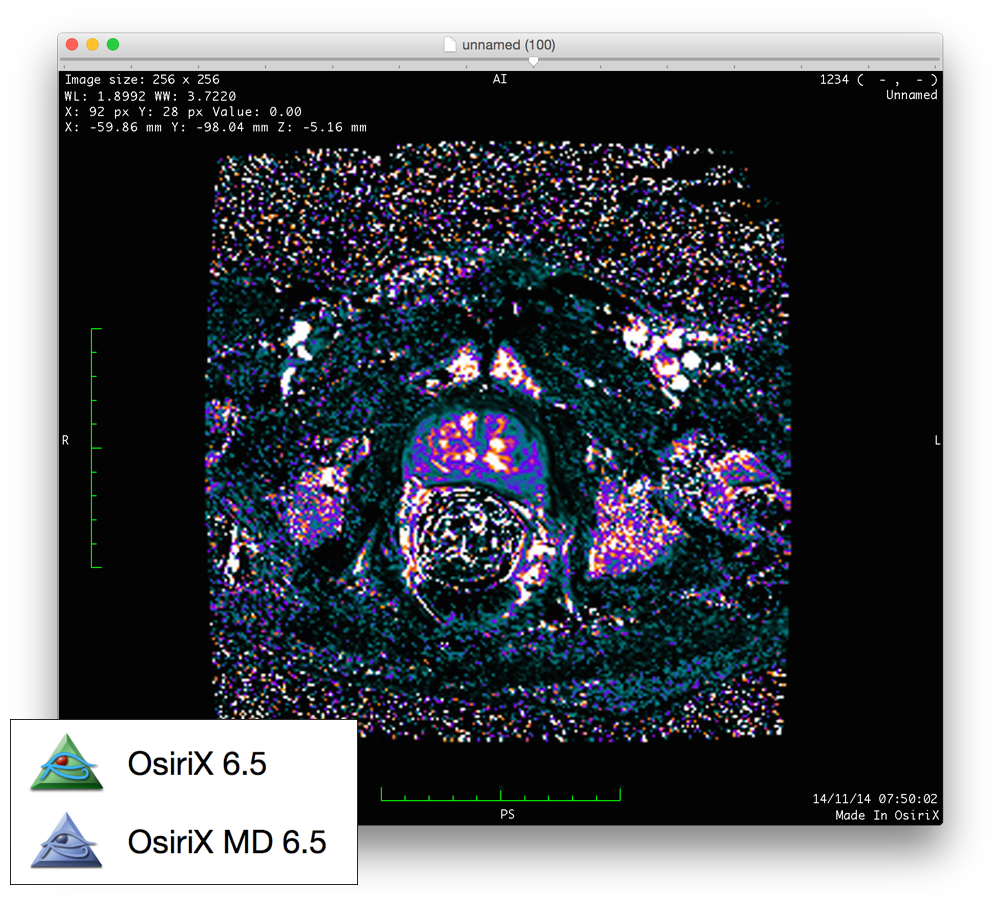
Unfortunately, DICOM Viewer for Mac is not available, but there are other tools that will help you view your DICOM images on Mac. This programme is only available on Apple computers, hence why so many radiologists own MacBooks.ĭICOM Viewer by CoreWare is an application that allows users to view DICOM images (images used for computer radiography).
#Osirix lite nifti to dicom software#
This is a free open source version of the software used by the Royal College of Radiologists for the viva part of the Final FRCR 2B exam, so obviously it makes sense to use it for teaching as well. In the hospital environment, this forms part of the picture archiving and communication system (PACS), which doctors will be familiar with!Ī popular software for radiologists working in the UK is currently a programme called 'Horos'. DICOM viewing software allows radiology trainees and consultants to view and manipulate medical images (such as radiographs or MRI scans) on their own PC, laptop or tablet. It is the standard for handling, storing, printing, and transmitting information in medical imaging. Parser.DICOM stands for 'Digital Imaging and Communications in Medicine'. add_argument( "-grabMetadata", action = "store_true", dest = "grabMeta", default = False) add_argument( "-configFile", dest = "json_config", required = True) add_argument( "-niftiDir", dest = "output_dir", required = False) add_argument( "-tablePath", dest = "table_dir", required = False) add_argument( "-dicomDir", dest = "input_dir", required = False) ArgumentParser( description = 'HorosExportToNiftiForHCC') join( output_dir, row, "")) = False:Ĭonvert_to_nifti( "arterial", art_row, output_dir)Ĭonvert_to_nifti( "venous", ven_row, output_dir) join( output_dir, row, "")) = False and os. Print ( "After Sort " + str( unique_times)) Print ( "Before Sort " + str( unique_times)) Print ( "Splitting Folder By Acquisition Time") Print ( "Error with dcm2nii for " + row + row + ", skipping")ĭef split_then_convert( image, row, output_dir, config): Print ( "Writing to this dest filename " + os. Print ( "Converting from this source dir " + os. Print ( "Already converted, or inappropriate tag " + str( row))ĭef convert_to_nifti( name, row, output_dir): Split_then_convert( image, row, output_dir, config)Ĭonvert_to_nifti( image, row, output_dir) join( output_dir, acc, image + ".nii.gz")) = True: Tagged_row = ( row for row in reader if row in )
Print ( "Starting Conversion with " + str( config) + " Configuration") insert( 0,)ĭef write_dicom_table( dicom_table, table_dir):ĭef write_by_series( json_config, input_table, output_dir):
#Osirix lite nifti to dicom series#
#Sort by MRN, then ACC, then Acq Time, then Series Numberīig_table = sorted( big_table, key = itemgetter( 1, 4, 5))īig_table. #print ("Key Error for "+acc+series_desc)
#print (mrn,acc,os.path.join(input_dir,i,j,k),acquisition_time,series_desc)
join( slugged_series_folder, dicoms_in_series))īig_table. join( slugged_series_folder, dicoms_in_series), force = True) move( series_folder_path, slugged_series_folder)ĭicoms_in_series = os. move( study_folder, slugged_study_folder)įor k in os. If config = "CTA-2phase":ĭef make_big_table( input_dir, json_config): Print ( "****Sorted through possible crap in " + i) #make_folder_if_absent(os.path.join(output_dir,i))įor j in os. append( arr, arr, axis = 0)ĭef check_directory( input_dir, json_config):įor tag, series in enumerate( config): Print ( "Confirm Spacing of New Flip-Only Routine") # Find the files that matches the given pattermįor filename in fnmatch. dcm2nii import Dcm2niixįor rootDir, subdirs, filenames in os.
#Osirix lite nifti to dicom code#
# 2) After desired tags are annotated in the "big table" manually (or in the future a more clever way) the code is run again reading this annotated "big table" and converts tagged series acccordingly into nifits and puts then in a folder named by accession numberįrom nipype. User then assigns a tag in the column following series description that will correspond to an abbreviated output file name for a converted nifti. # 1) A csv "big table" is created with a row for each series of each study. # This takes DICOM files of studies with multiple series exported from Osirix (with hierarchy and non-deidentified options selected) and converts them into Nifti files in a two part process:
